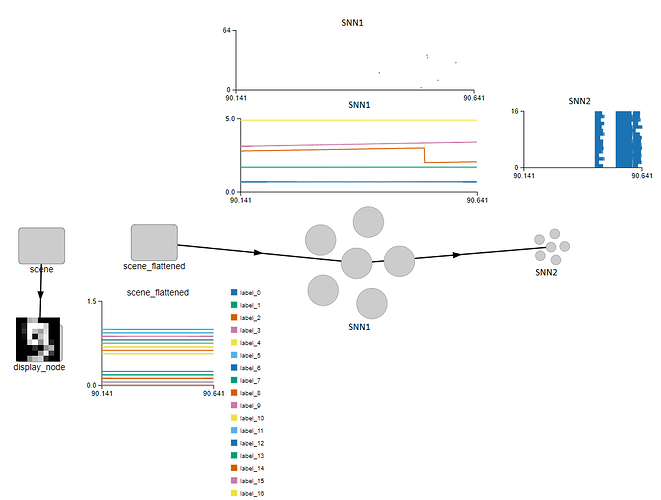
My goal is just to build at least a two-layer SNN with connections initialized randomly and then trained using NengoDL on the 8x8 handwritten digit MNIST. After training, i will use the connection weights to hardcode the connections in a circuit. I may have to use decoders for efficiency so i think I should try to understand how it works and why.
For this specific use case, because the connections between the two layers in the SNN are randomized, that connection won’t use decoders and encoders. As mentioned in a previous post, encoders and decoders can be combined to form the connection weights. Thus, you can reverse that formulation and “derive” the encoders and decoders from a set of trained connection weights by factoring the weights into their appropriate components (this is typically not easy to do though, so connection weights trained in this way are typically left unfactored). In your example, the one place you might need to use decoders on the readout layer (or the second layer’s output).
Why do we want to map back the neural activity into the N-dimensional input space? If we want to compute f(x) = x^2 for instance, wouldnt it be easier to compute that classically before feeding into the neural network?
The NEF (Neural Engineering Framework) was developed with the design philosophy of implementing as much of the computation in the spiking neural network, with the eventual goal of implementing entire systems (end-to-end) wholly within a spiking neural network. Spaun for example is a vision-to-cognition-to-motor system completely implemented in a spiking neural network. For such networks, the ability to compute decoders to absolutely crucial since training a network of that size from scratch is close to impossible. For hardware implementations the NEF gives us the ability to implement arbitrary computations in SNN so that we can take full advantage that SNN in hardware provides.
However, if you do not have these restrictions in mind with your network, then using the NEF to compute decoders that approximate any function is not necessary.
If we want to compute a function that is hard to define, say a function that performs edge detection on the input how are these decoder weights specified? I should specify the function it wants to compute?, f(x) = x or some other function?
The beauty of Nengo is that you can give any arbitrary python function to the decoder solver when creating a connection. What Nengo does is to use this python function to create a mapping between a set of inputs (randomly generated) to the corresponding set of outputs (determined by the logic of the given function). This mapping is then used to solve for the ensemble’s decoders. This method can be used to approximate the edge detection functionality, but I haven’t tried it myself, so I can’t comment on the effectiveness of such an approach.
For more complex functions, it is typically more helpful to break down the function into simpler components. For your example of edge detection, you could try applying a filter (e.g., a Gabor filter - see Nengo example here), and then performing a function on the output of the filters (i.e., the feature vector).
When training the SNN, what happens to these decoder and encoders?
In NengoDL, you can specify which Nengo object to train. Only objects with a trainable=True attribute will have their properties modified during the training process.
Also, say I have an input 8x8-pixel Image as an input to an SNN. I understand I cant use this directly, so I flattened it to a 1x64 vector. The first SNN layer has 64 neurons. I want the ith neuron to receive as its single input, only the ith pixel in the 1x64 input vector. It’s a one-to-one connection. what would be the parameters in nengo.connection
You can achieve this by modifying the encoders of the ensemble. Here’s how you would do it:
import nengo
import numpy as np
from nengo.dists import Samples
d = 64
n = d
eye = np.eye(d)
with nengo.Network() as model:
ens = nengo.Ensemble(n, d, encoders=Samples(eye))
# Note, you can use the `Choice` distribution here as well. `Sample` chooses the rows in order (it cycles when it reaches the end of the list), while `Choice` chooses the rows using a normal distribution.
I should note that the bias and gains for each neuron in the ensemble are chosen at random. If want to fix the neuron response curves to make them all identical, you should do:
ens = nengo.Ensemble(d, d, encoders=Samples(eye), intercepts=Choice([0]),
max_rates=Choice([200]))
Alternatively, I wanted to modify the currents on each neuron in the SNN layer directly, but I’m not sure how to do that.
Can you elaborate on what you want to do with the neuron currents? Do you want to inject a constant bias, or to modify the currents on the fly while the simulation is running?