Hi! Perhaps I can help you with your questions:
====================
I am trying to build a bionic network. How to realize the excitatory and inhibitory connection between neurons?
This really depends on specific constraints in the excitatory & inhibitory connection implementation. If your primary concern is with building a network that transfers information from population A’s neurons to population B’s neurons without wanting to worry about setting up the individual excitatory and inhibitory connections, then Nengo already does this for you.
pop_A = nengo.Ensemble(n_neurons=10, dimensions=1)
pop_B = nengo.Ensemble(n_neurons=20, dimensions=1)
nengo.Connection(pop_A, pop_B)
And that’s it! With the code above, Nengo will create a connection between population A and B and solve for the appropriate connection weights in order to perform the x=y function. Note that in the code above, the connection weights that Nengo solves for you contains a mix excitatory and inhibitory connections.
If you are looking to create a network where the excitatory and inhibitory connections are separate connections, you can use something like the nengo-bio codebase (as @zerone mentioned), or do what we refer to as the “Parisien” transform. This transform is used to convert a “standard” Nengo connection, where both inhibitory and excitatory connections can originate from the same neuron, into a network where inhibitory and excitatory connections are divided so that an individual neuron can either only be inhibitory or excitatory.
There is a fairly large github thread discussing it, with an example Nengo model defined here. Not that this uses the Dales solver from the parisien branch of the nengo-extras repository. The Parisien transform can also be done using nengo-bio with example code here.
I’m not entirely sure what your use case is, but you did mention NengoDL in your original query? If you are using NengoDL, some work will be needed to convert these example for use in NengoDL.
====================
The weights are all set to 1, I think the B neuron will completely convey the A neuron’s signal. But this is not the case.
What you are observing may be a result of a slight misunderstanding of what information Nengo is showing to you. When you do something like this:
pop_A = nengo.Ensemble(50, 1)
probe_A = nengo.Probe(pop_A, synapse=0.005)
what probe_A is probing is the decoded output of the ensemble pop_A. In order to get the decoded output of an ensemble, Nengo performs an optimization calculation to determine a set of output weights such that for a specific set of inputs (to the neural ensemble), the decoded output (i.e., the spike rate multiplied by the output weights) of the neural ensemble performs a specific function. By default, the function is an identity function (i.e., output = input). However, because Nengo generates neurons with randomly determined response curves, this often means that the output weights for any specific neural ensemble contain both positive and negative numbers. Going back to your question, where you specified a neurons-to-neurons connection weight of all 1’s, what this means is that your A population is indeed driving all the neurons in the B population, but because of the mix of positive and negative output weights, the decoded output averages out to 0. To see the effect of population A driving the neurons of population B, you’ll need to probe the spikes of the B population:
probe_spikes_B = nengo.Probe(pop_B.neurons)
====================
How to set negative weights? If possible, can you provide some examples for me to learn from? Thank you very much!
You’ve almost got it in the original example code you provided. Creating an inhibitory connection in Nengo works something like this:
pop_A = nengo.Ensemble(50, 1)
inhibitory_input = nengo.Node()
inhibitory_conn = nengo.Connection(inhibitory_input, pop_A.neurons,
transform=[[-1]] * pop_A.n_neurons)
If connecting from another neural ensemble:
pop_A = nengo.Ensemble(50, 1)
pop_B = nengo.Node()
inhibitory_conn = nengo.Connection(pop_B, pop_A.neurons,
transform=[[-1]] * pop_A.n_neurons)
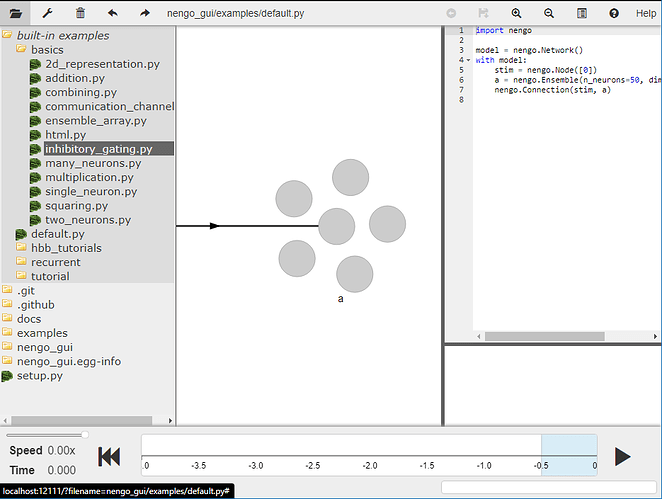
There is also an example of creating inhibitory connections in our NengoGUI. You can find it under “built-in examples” → “basics” → inhibitory_gating.py
If you don’t have NengoGUI installed, you can find instructions on how to do so here.