Is there a way to remove (or set to zero) specific synapses? Given an ensemble A that outputs to ensemble B, I’d like to lesion the network by choosing some subset of neurons in B and removing all incoming connections to those neurons coming from A, while still having other connections to those neurons from other ensembles or nodes. I know how to zero a neuron’s activations from my previous question, but I would now like to target specific synapses. If anybody knows how to do this or can point me to somewhere in the Nengo codebase where synapses are exposed that would be great!
Hi @Luciano,
You can “lesion” specific synapses in a Nengo network using a similar method to the neuron lesion code you linked in your post. First, it is important to note that in Nengo, a synapse is a filter applied to the connection between two ensembles. The synaptic filter is applied on the collective input to a neuron on the post population. This differs slightly from the biological definition of a synapse, which is between individual neurons. Thus, in the method I will outline below, to achieve the “lesioning” you desire, we modify the weights of the connection between the two ensembles, rather than modifying the synapse itself.
Similar to the “ablate neurons” function, a “lesion connection” function can be defined as follows:
def lesion_connection(sim, conn, lesion_idx):
connweights_sig = sim.signals[sim.model.sig[conn]["weights"]]
connweights_sig.setflags(write=True)
connweights_sig[lesion_idx, :] = 0
connweights_sig.setflags(write=False)
Note that the lesioning index code is dependent on how the nengo.Connection is created. If a connection between two ensembles is created like so (the default way):
conn = nengo.Connection(ensA, ensB)
The weights signal (i.e., sim.signals[sim.model.sig[conn]["weights"]]) has a shape that is (1, ensA.n_neurons). This means that it only contains the decoders of ensA. If you want to lesion the output of a neuron in ensA to every neuron in ensB it is connected to, no additional modification to the nengo.Connection is needed, and the lesion function can be used as is.
However, since you want to achieve the reverse (lesion all inputs to a specific neuron in ensB), we’ll need to change the code slightly. Namely, when we create the nengo.Connection, we specify the solvers with the weights=True flag to force the connection to be created with the full weight matrix. This weight matrix combines the decoders of ensA with the encoders of ensB. The full code for this is as such:
conn = nengo.Connection(ens1, ens2, solver=nengo.solvers.LstsqL2(weights=True))
With this change to the code, you can lesion the connection similar to how it was done with the neuron ablation code:
with nengo.Network() as model:
... # define your model
with nengo.Simulator(model) as sim:
lesion_connection(sim, conn, <lesion_index>)
sim.run(<runtime>)
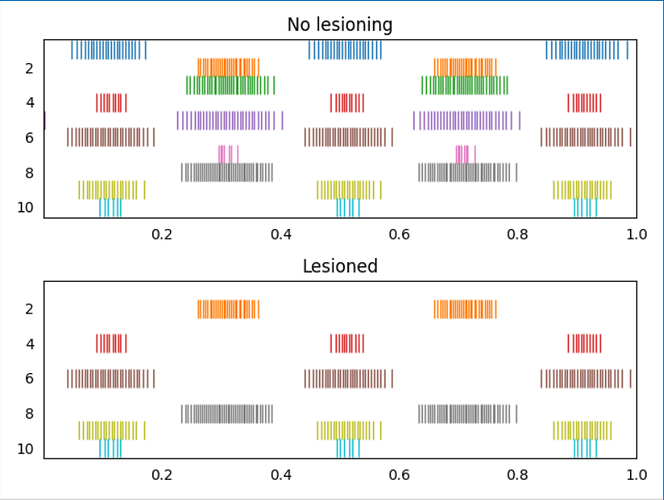
I’ve attached an example script (test_lesion_conn.py (1.5 KB)) that demonstrates this code. In the script, two ensembles are constructed, and here is the output plot of the script showing on non-lesioned run, and one lesioned run (the input connections to the 1st, 3rd, 5th, and 7th neurons in ens2 have been lesioned)
Some additional notes:
- The network is created with a seed so that multiple runs should be identical, which is why you see identical spike patterns for the non-lesioned connections.
- The lesioning is applied on a per-connection basis, so, you should be able to achieve the desired functionality of leaving other connections to “ens B” intact. You can test this out in the example code by adding an additional ensemble and connection to
ens2.
Is this possible with NengoDL? I’m getting AttributeError: 'Simulator' object has no attribute 'signals'.
It may be possible to do the synapse removal in NengoDL, but it depends on the specifics of your model. Can you provide some example code so that I can investigate how it would work with your code?
I’m using this code, specifically trying to run a03_GC_PMC_line.py. In framework.py I replaced:
conn = nengo.Connection(net.error[:dim], net.M1.input[dim:])
with
net.pmc_m1_conn = nengo.Connection(net.error[:dim], net.M1.M1[dim:], solver=nengo.solvers.LstsqL2(weights=True))
and in a03_GC_PMC_line.py I added:
def on_start(sim):
ablate_synapses(sim, net.pmc_m1_conn, range(9000))
Hmmmm. The original code I posted is written specifically for the core Nengo backend, and it achieves the synapse ablation through “hacky” means (because we are accessing very low level information within the nengo.Simulator object). So… if you want to use the NengoDL simulator to run the model, it’ll have to be changed to support that backend. I’m not 100% sure if such a functionality is even supported with NengoDL, although from my initial investigation of the code, I don’t think it’s possible. But, I’ll keep you posted! 
In the mean time, if you do want to use NengoDL to train your model, you can try training your network in NengoDL (without the ablation code), then using the nengo_dl.Simulator.freeze_params() functionality to convert the trained NengoDL model back into a Nengo model. Once you have the standard Nengo model, you can then run your Nengo simulation with nengo.sim and the ablation code.
I wanted to add another tensorflow net that would stimulate M1, learning an optimal stimulus based on resulting arm movement. This could learn to compensate for the ablation, the idea is that of the neural coprocessor. I don’t think I could accomplish this by freezing the params since the training of the tensorflow net depends on the simulation of the rest of the (ablated) model. I’ll look more into the source code, thanks again for your help!
Just to get a better idea of your workflow, so that I may suggest other potential approaches, am I correct to understand that you are attempting to do this:
- Create a model in Nengo
- Apply the ablation to the model
- Add a tensorflow (or nengo-dl) network on top of the model
- Train in NengoDL (or TF)
Is there an additional simulation step between 1 and 2?
Are there multiple back and forth simulations between regular Nengo and NengoDL?
Essentially what I’m trying to do is:
- Create a model in Nengo
- Record neural activity in M1 and arm position during an arm reaching task
- Train a tensorflow network called EN using that recorded data (outside of Nengo)
- Integrate EN to the Nengo model, it now predicts arm movement based on M1 activity during the reaching task (in NengoDL)
- Ablate some PMC -> M1 synapses (apparently not possible in NengoDL)
I’m here - Add a tensorflow network called CPN which will predict M1 activity based on PMC. CPN outputs to EN and is trained through backpropped error from EN network to learn the optimal M1 activity (stimulus) to drive the arm towards the target. This requires NengoDL because I want to train CPN and have it stimulate M1 in realtime.
It may also be possible for me to achieve step 6 by first recording PMC and M1 activity for the Nengo model with ablated synapses, training CPN outside of the simulation, then returning to the simulation with both networks trained and using the strategy of freezing the parameters to use the CPN to stimulate M1 using Nengo.
I messed around with Nengo and NengoDL a bit and I believe I have found an approach that will work with your workflow. In essence, this approach is to utilize Nengo’s ability to specify the full connection weight matrix for a connection; i.e., like so:
conn = nengo.Connection(ens1.neurons, ens2.neurons, transform=weights)
and use this to perform the connection ablation. The idea is as follows:
- We use the Nengo (or NengoDL) simulator to solve for the “optimal” connection weights for us, as per usual.
- Extract the solved connection weights from the Nengo simulator object.
- Perform the appropriate ablation on the solved connection weights.
- Recreate the model, but use the neuron-to-neuron connection to create a connection with the ablated weights.
- Since the recreated model is a standard Nengo model, and the ablated weights are defined in the model, rather than having to mess with the simulator signals, we should be able to simulate the model with NengoDL with no problems.
Now, if your model is particularly big, it might take a while to build the whole model, and this would be inefficient if your goal is to build the model just to extract the initial (optimal) connection weights. To get around this, we can define a function that creates a subnetwork with just the components involved in the ablated connection. Then we can create a Nengo simulator object for this subnetwork only, reducing the overall build time.
I’ve implemented this approach in this example code (which builds off the previous example code): test_lesion_conn2.py (5.1 KB)
Note that this neuron-to-neuron connection approach only works with ablating connections. A similar, but more involved approach is needed if you want to lesion specific neurons as well.
Awesome, thanks! I’m going to try to adapt this to my application and I’ll let you know if I have any questions.
I refactored my code to use this approach but now I’m getting the following output:
Warning: Simulators cannot be manually run inside nengo_gui (line 83)
If you are running a Nengo script from a tutorial, this may be a tutorial
that should be run in a Python interpreter, rather than in the Nengo GUI.
See https://nengo.github.io/users.html for more details.
must declare a nengo.Network called "model"
I’m trying to run in nengo gui. The warning is happening when I create the nengo_dl simulator to create the ablated weights (line 103 of your code). I also tried using the on_start(sim) hook to build the weights using the nengo gui simulator, but the problem with that is I need the weights in the generate function to create the subnet, but on_start executes after generate. Do you know if there’s a way to access the nengo gui simulator in or before generate, or do you know some other way to do this? If you want to look at the code I can add you to my github repo.
Ah, I did not realize you were using NengoGUI to run your code. Invoking additional simulators in NengoGUI is a tricky thing to do, because NengoGUI hijacks the simulator call to insert its own simulator to run the simulation. While it has been achieved in the past, the approach used to run another Nengo simulation in NengoGUI is hacky at best, and not officially supported.
So… I think your best approach is to refactor your code and pull out the weights generation network into a separate script that you can run outside of NengoGUI (just run it in a terminal with python <script_name.py>). In that script, you can use NumPy’s save features to save the weights to file. Run this network to generate the weights before you run your actual network in NengoGUI, and in the NengoGUI network, use NumPy’s load function to load the saved weights from file.
As an example, this code saves the ablated weights to file:
build_net = create_subnet(seed=0)
with nengo_dl.Simulator(build_net) as build_sim:
new_weights = make_ablated_weights(build_sim, build_net.conn, ablate_idx)
np.savez_compressed("ablated_weights.npz", weights=new_weights)
And then this code loads it from file:
with nengo.Network() as model2:
new_weights = np.load("ablated_weights.npz")["weights"]
subnet = create_subnet(seed=0, weights=new_weights)
Here’s some example code showing it running (albeit in one script, but the idea is there): test_lesion_conn3.py (5.2 KB)
Ok hopefully this is the last issue  : in my network, the connection I’m trying to ablate is from an ensemble of representational dimension 2 to one of dimension 4, so my connection is from
: in my network, the connection I’m trying to ablate is from an ensemble of representational dimension 2 to one of dimension 4, so my connection is from net.ens1 to just the first 2 dimensions of net.ens2, which is net.ens2[:2], but when I do this I get this KeyError:
File "test_lesion_conn3.py", line 93, in make_ablated_weights
conn_weights, np.matrix(sim.model.params[conn.post].gain).T
KeyError: <Ensemble (unlabeled) at 0x7fd99e40cd90>[:2]
Here’s an edited version of your script which replicates the issue: test_lesion_conn4.py (5.3 KB)
Ah, yes. My original implementation was not “slice-safe”. The fix is simple though, instead of using conn.post, you need to use conn.post_obj. Here’s my version of the script that should work with slicing. In my script I also added a common input function to use across both models.
test_lesion_conn4.py (5.3 KB)
To clarify, the reason we can’t just do something like the following (in nengo gui):
def on_start(sim):
new_weights = make_ablated_weights(sim, net.conn, ablate_idx)
net.conn = nengo.Connection(
net.ens1.neurons, net.ens2.neurons, transform=new_weights
)
is because we can’t change the connection at runtime right?
Yes, sort of. When you do something like this (NengoGUI automatically does this when you press the “play” button):
with nengo.Simulator(model) as sim:
sim.run(1)
a few things are actually happening.
- First, the Nengo model (
model) is “compiled” (connection weights are solved for, mapping between ensembles are done, etc.,) and converted into a bunch of connected signals and operators that, when put together, define the behaviour of your model. - Next, all of these objects are stored with the
sim, as well as placeholders for simulation data, probes, etc. - Finally, the
runfunction is called, and this does the actual simulation and populates the probe data and what not.
Any function defined in on_start (for NengoGUI) is run after the simulator object is created, and before the run function is called (i.e., between steps 2 and 3). At this point in the build process, all of the connection weights have already been defined and the Nengo model has already been compiled down into the equivalent operators and signals, so defining a nengo.Connection here has no effect because the build process is not re-initiated after the on_start call is made.
So, the specific on_start function you defined won’t work as you intended, yes, but because at the time it is called, the Nengo model has already been compiled, and further changes to the original model are not reflected in the compiled model stored in the sim object (because the “compiler” [we call it a “builder” in the codebase] is not called any more).
As an aside, you can technically change the connection weights at runtime (using a method similar to the original lesion_connection function. I.e., directly modifying the sim.signals values), but this is a rather “hacky” approach and not recommended. ![]()