Hi,
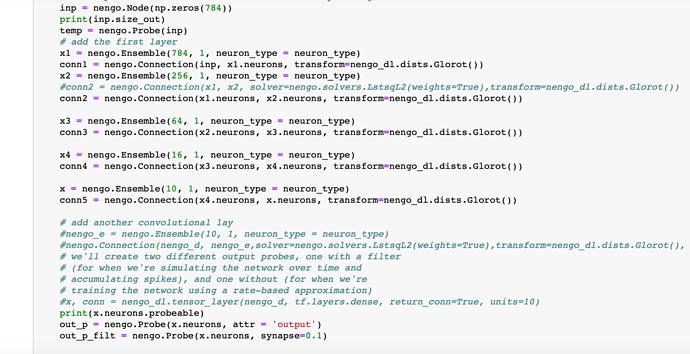
Please find the code pasted below. We have run the code in jupyter notebook, I have divided the code using “********” for blocks that have been run in different cells. As mentioned the final error after training is not consistent. Please let us know where the issue is.
note: changed the comment to // as using # is causing format issues.
%matplotlib inline
import gzip
import pickle
from urllib.request import urlretrieve
import zipfile
import nengo
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import nengo_dl
urlretrieve(“http://deeplearning.net/data/mnist/mnist.pkl.gz”,
“mnist.pkl.gz”)
with gzip.open(“mnist.pkl.gz”) as f:
train_data, _, test_data = pickle.load(f, encoding=“latin1”)
train_data = list(train_data)
test_data = list(test_data)
print(len(test_data[1]))
for data in (train_data, test_data):
one_hot = np.zeros((data[0].shape[0], 10))
one_hot[np.arange(data[0].shape[0]), data[1]] = 1
data[1] = one_hot
n_in = 784
n_hidden = 64
minibatch_size = 50
with nengo.Network() as auto_net:
# input
nengo_a = nengo.Node(np.zeros(n_in))
print(nengo_a.size_out)
# first layer
nengo_b = nengo.Ensemble(
n_hidden, 1, neuron_type=nengo.RectifiedLinear())
print(nengo_b.size_in)
nengo.Connection(
nengo_a, nengo_b.neurons, transform=nengo_dl.dists.Glorot())
# second layer
nengo_c = nengo.Ensemble(
n_in, 1, neuron_type=nengo.RectifiedLinear())
nengo.Connection(
nengo_b.neurons, nengo_c.neurons,
transform=nengo_dl.dists.Glorot())
# probes are used to collect data from the network
p_c = nengo.Probe(nengo_c.neurons)
with nengo.Network() as net:
# set some default parameters for the neurons that will make
# the training progress more smoothly
net.config[nengo.Ensemble].max_rates = nengo.dists.Choice([100])
net.config[nengo.Ensemble].intercepts = nengo.dists.Choice([0])
neuron_type = nengo.LIF(amplitude=0.01)
nengo_dl.configure_settings(trainable=True)
# the input node that will be used to feed in input images
inp = nengo.Node(np.zeros(784))
print(inp.size_out)
temp = nengo.Probe(inp)
# add the first layer
x, conn1 = nengo_dl.tensor_layer(inp, neuron_type, transform=nengo_dl.dists.Glorot(),shape_in=(256,), return_conn=True)
x, conn2 = nengo_dl.tensor_layer(x, neuron_type, transform=nengo_dl.dists.Glorot(),shape_in=(64,), return_conn=True)
print(conn2.size_out)
x, conn3 = nengo_dl.tensor_layer(x, neuron_type, transform=nengo_dl.dists.Glorot(),shape_in=(16,), return_conn=True)
x, conn4 = nengo_dl.tensor_layer(x, neuron_type, transform=nengo_dl.dists.Glorot(),shape_in=(10,), return_conn=True)
#x, conn5 = nengo_dl.tensor_layer(x,neuron_type,transform=nengo_dl.dists.Glorot(), shape_in=(10, ), return_conn=True)
print(x.probeable)
out_p = nengo.Probe(x)
out_p_filt = nengo.Probe(x, synapse=0.1)
minibatch_size = 200
sim = nengo_dl.Simulator(net, minibatch_size=minibatch_size)
// add the single timestep to the training data
train_data = {inp: train_data[0][:, None, :],
out_p: train_data[1][:, None, :]}
// when testing our network with spiking neurons we will need to run it
// over time, so we repeat the input/target data for a number of
// timesteps. we’re also going to reduce the number of test images, just
// to speed up this example.
n_steps = 30
test_data = {
inp: np.tile(test_data[0][:minibatch_size2, None, :],
(1, n_steps, 1)),
out_p_filt: np.tile(test_data[1][:minibatch_size2, None, :],
(1, n_steps, 1))}
def objective(outputs, targets):
return tf.nn.softmax_cross_entropy_with_logits_v2(
logits=outputs, labels=targets)
opt = tf.train.RMSPropOptimizer(learning_rate=0.001)
def classification_error(outputs, targets):
return 100 * tf.reduce_mean(
tf.cast(tf.not_equal(tf.argmax(outputs[:, -1], axis=-1),
tf.argmax(targets[:, -1], axis=-1)),
tf.float32))
print(“error before training: %.2f%%” % sim.loss(
test_data, {out_p_filt: classification_error}))
do_training = True
#summ_list = []
if do_training:
# run training
sim.train(train_data, opt, objective={out_p: objective},n_epochs=1)
sim.save_params("./mnist_params")
print(“error after training: %.2f%%” % sim.loss(
test_data, {out_p_filt: classification_error}))