Hi,
I’m trying to make a few changes to the Vision example for mnist.
To make it a little more … CNN-like
My plan is to try and make a SCNN that can work with single spike instances, then eventually work with a spike-timing learning rule. My alternative approach seems to be hand-coding all the individual pixels to do this, but i imagine there must be a more efficient way
i change the Gabor to a predefined 5x5 kernel and i am only testing it on one feature just now.
i also change the receptive field locations from randomly sparse to sequential with a stride of one.
So i should have 576 output from this, which i do 
but when it comes to taking the values out of the Mask.populate function is when i am getting confused
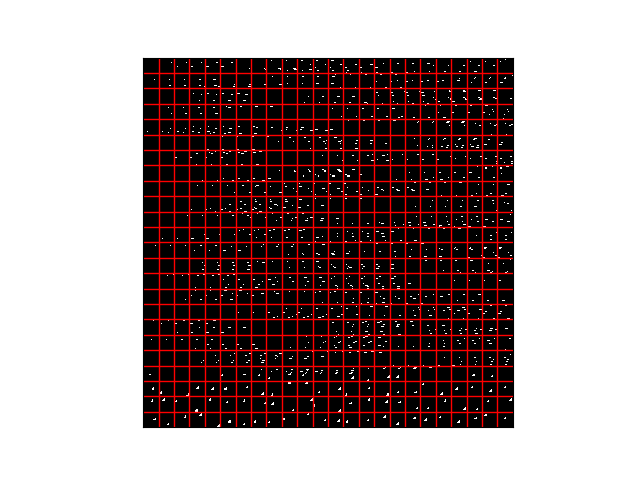
i get a 576,784 tuple which is cool 576 28x28s flattened
but when i try to tile to see what the representation of this is the kernels are now randomly spread across different masks 
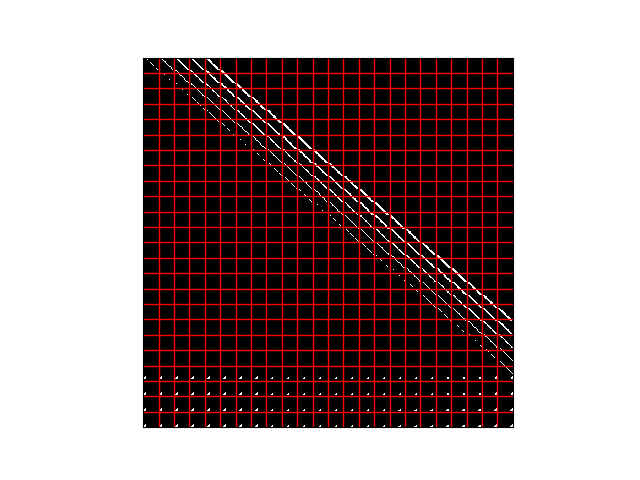
the bottom few rows almost look good but the rest is just confusing…
?? Is this just the tiling program and all the reshaping making it look odd, or is this what the neurons i have ensembled are doing ??

