Hi @myyim, welcome to the forum!
A Nengo model can look noisy in lots of ways that are still deterministic. In your model, for example, the only random elements are the neuron properties, which could be made deterministic by setting an integer seed on the ensemble.
Your model looks noisy because a LIF neuron is a spiking neuron, which means that it approximates values by spiking. That means that when you look at the output of the ensemble as a whole, it’s always going to be, to a certain extent, “noisy” looking because you’re trying to figure out what’s happening at one moment in time which might be when a cell is spiking, or might be right after it’s spiked. We smooth things out with filtering so that we have some estimate of what’s going on, but it’s always a noisy estimate. The synapse argument defines those filters; probing a spiking ensemble with synapse=None is always going to be very very noisy.
The above explains why any value is never exactly that value. However, in your model, you also have no input, so the value should just be 0, which can be easily represented by the cells not spiking. Well, there is an additional detail, which is that we also inject a background bias current to mimic the background firing rate of biological cells receiving no synaptic input. Again, this is completely deterministic, but gives the appearance of noise despite there being no input.
Fortunately, if you don’t want to deal with the oddities of spiking neurons, you can simply use a rate-based neuron model. Try this:
Model = nengo.Network(label="Network")
with Model:
L0 = nengo.Ensemble( 2, dimensions=2,noise=None, seed=None, neuron_type=nengo.LIFRate())
probe = nengo.Probe(L0, synapse=None)
sim = nengo.Simulator(Model,0.001)
sim.run(1)
x = sim.data[probe]
You will now see that the output, x, is always the same. It is still non-zero, because we still inject a bias current, and since there are only 2 neurons in your ensemble, the bias makes the representation significantly far from [0, 0]. You can increase the number of neurons to keep the representation closer to [0, 0]:
Model = nengo.Network(label="Network")
with Model:
L0 = nengo.Ensemble(100, dimensions=2,noise=None, seed=None, neuron_type=nengo.LIFRate())
probe = nengo.Probe(L0, synapse=None)
sim = nengo.Simulator(Model,0.001)
sim.run(1)
x = sim.data[probe]
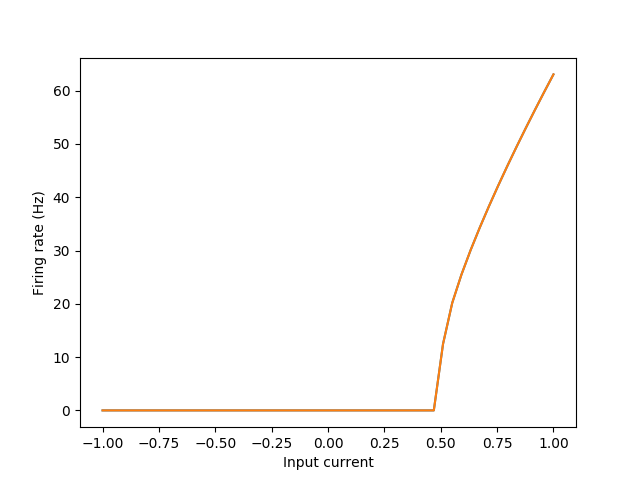
You can also play around with the neuron properties such that they do not receive bias current, or only negative bias current. See the tuning curves example for details on that.
Also, as @Seanny123 mentioned, you can also use the nengo.Direct() neuron type to bypass any kind of neural encoding / decoding, and instead use Nengo as a general dynamical systems simulator:
Model = nengo.Network(label="Network")
with Model:
L0 = nengo.Ensemble( 2, dimensions=2,noise=None, seed=None, neuron_type=nengo.Direct())
probe = nengo.Probe(L0, synapse=None)
sim = nengo.Simulator(Model,0.001)
sim.run(1)
x = sim.data[probe]
In this case, you will see that x is always exactly [0, 0].