Hi @n299! You packed quite a few questions in your post, so below I’ll try to address them individually (and slightly out of order, but hopefully it’ll make sense to you).
I’m looking at Nengo learning rules to figure out where the update is happening …
It’s awesome that you are looking at the Nengo code base to learn how it is doing things. Here’s a quick rundown of what’s happening when you specify a learning rule and build a nengo.Simulator object:
- Nengo iterates through all of the objects in your model and runs each one through the Nengo builder (
nengo.builder.builder.Model.build(...))
- If you have specified a learning rule on a connection in your model, it calls the
build method on the learning rule.
- For each learning rule type defined in Nengo, there are associated registered builder functions. It is this function that gets called by the
build method from the previous step. There is also a registered builder function generic to all learning rules.
- The “job” of each builder function is to construct an operator graph and add it to existing operator graph for the Nengo model. For learning rules, you’ll notice that most of them add a
Sim<Name> operator. These operators are in turned defined in builder.learning_rules.
- When an operator is added to the Nengo model, it’s
make_step function is called to determine what computation should be added.
- Once the
make_step function has been processed, the builder continues on to the next Nengo object in the model.
Right, with that “quick” overview done, let’s get to answering your questions! 
==========
I’m looking at Nengo learning rules to figure out where the update is happening, and it seems to be in the “step_sim” function, but I want to know what “post_filtered” and “pre_filtered” mean.
Also, is what is being returned in the below rule what is being added to the weight matrix
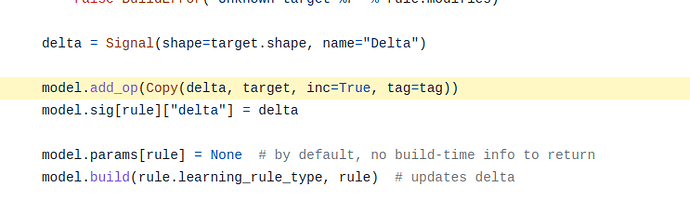
Well, sort of. For learning rules, the update is sort of happening in multiple places. The code to calculate the exact change in values (I’m being expressly vague here and not saying “change in weights” for a reason) caused by the learning rule is done in the step_<name> function. You are correct in that respect. Using the step_simoja function as an example, the value changes are applied to the delta nengo.Signal object. This signal itself is created in the generic learning rule builder function, and passed to the SimOja class when it is created.
So, the step_simoja call updates the value of the delta signal, but this has not yet updated any weights in the model. The weight updates are done by an operator added by the generic learning rule builder, where target is defined according to what the learning rule affects.
To summarize, the updates for a learning rule are performed in two steps:
- The
step_sim<name> function (from the Sim<Name> operator) computes the change in values and updates the delta signal.
- The
Copy operator copies the value of the delta signal to the target signal (which happens to be the connection weights for the Oja rule.
==========
but I want to know what “post_filtered” and “pre_filtered” mean
Documentation for the SimOja learning rule operator can be found here. To answer your specific question about pre/post_filtered, these variables are the nengo.Signals that represent, respectively, the spike trains from the pre and post ensembles of the learning rule, filtered by any synapses defined for these spike trains.
==========
Is there a way to access the “spiked” values of two layers being connected via a learning rule so that I can use them to update the weights?
Yes! If you define the Oja learning rule with synapse values of None, this will leave the pre and post spike trains unfiltered, so you get the raw spiking data. You can do this with:
nengo.Connection(pre, post,
learning_rule_type=Oja(..., pre_synapse=None, post_synapse=None))
Note that the spike data will still be accessible to the learning rule step function through pre/post_filtered. There is no special variable used to denote non-filtered data specifically. Additionally, the spike data will be in a format where spikes are represented with a value that is 1/dt. This is done to keep the area of a single spike to 1 (1/dt * dt = 1).
==========
Also, for the below rule, where is alpha being determined? Is it when you call the rule when you build a model?
For the Oja rule in particular, alpha is computed when the make_step function is called (see Step 5 in the quick rundown above). And you are correct, this is done when nengo.Simulator(model) is called to build your Nengo model.