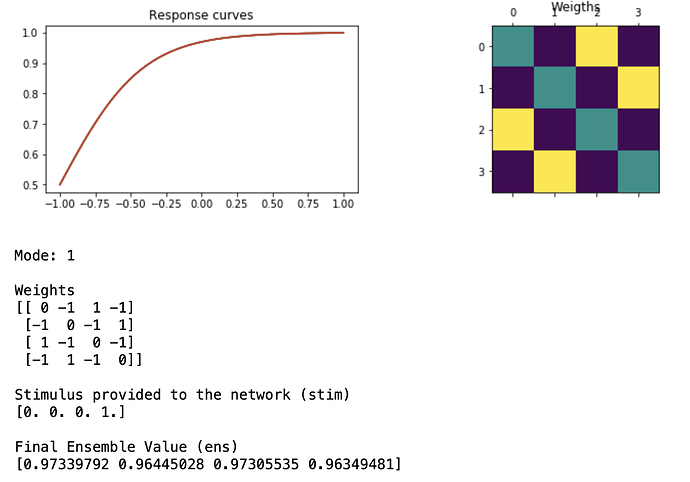
I’ve started working on implementing something close to a Hopfield NN in Nengo and was struggling with a few points:
-
I was thinking of representing my inputs as n-dimensional vectors whose elements are 0 or 1.
Is this a good idea?
Is the best way to do this by using something likeEnsembleArray(n_neurons=10,n_ensembles=dimensionality)? -
As I’m planning on using a custom Hebbian learning rule that takes the pre and post neural activities, I’d like to have direct access to these in my learning
Node()object.
I’ve previously achieved this withEnsemble()by:
pre_nrn = 10
post_nrn=10
pre = nengo.Ensemble(
n_neurons=pre_nrn,
dimensions=1,
encoders=generate_encoders( pre_nrn ),
# intercepts=[ 0.1 ] * pre_nrn,
label="Pre"
)
learn = nengo.Node(
output=memr_arr,
size_in=pre_nrn + post_nrn,
size_out=post_nrn,
label="Learn"
)
nengo.Connection( pre.neurons, learn[ :pre_nrn ], synapse=0.005 )
nengo.Connection( learn, post.neurons, synapse=None )
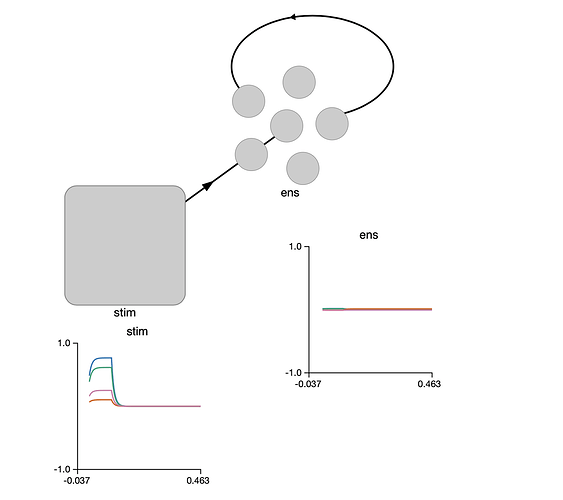
I’m currently trying the following to achieve an equivalent effect with an EnsembleArray() for an input vector of 3 elements, but I’m getting the error
nengo.exceptions.ValidationError: init: Shape of initial value () does not match expected shape (10, 30):
pre_nrn = 10
post_nrn=10
dimensionality = 3
pre = EnsembleArray(
n_neurons=pre_nrn,
n_ensembles=dimensionality,
label="Pre"
)
pre.add_neuron_output()
learn = nengo.Node(
output=memr_arr,
size_in=(pre_nrn + post_nrn) * dimensionality,
size_out=post_nrn * dimensionality,
label="Learn"
)
nengo.Connection( pre.neuron_output, learn[ :pre_nrn ], synapse=0.005 )
I would have expected pre.neuron_output to have shape (10,3), what am I doing wrong?
 )
)