Welcome to challenges!
The idea is to have a series of simple problems, presented every so often, where people can discuss different circuits and implementations / weigh the pros and cons of various approaches. The goal is to hopefully learn some new techniques for neural circuit design and generate a set of efficient, robust solutions to various problems.
Answers should be posted with code and simulation results!
The first challenge is to generate a circuit that transforms an input signal into a clean 0 if it’s below .5, and a clean 1 if it’s above .5.
The input to the system is nengo.Node(lambda x: np.cos(x)*.5+.5) and the system should run for 6 seconds.
Go!
1 Like
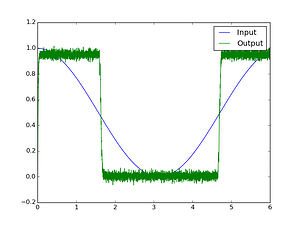
Here’s one possible solution:
import nengo
import numpy as np
model = nengo.Network()
with model:
stim = nengo.Node(lambda x: np.cos(x)*.5+.5)
b = nengo.Ensemble(n_neurons=50, dimensions=1,
intercepts=nengo.dists.Uniform(.5, 1),
encoders=[[-1]]*50)
out_b = nengo.Ensemble(n_neurons=50, dimensions=1)
output = nengo.Ensemble(n_neurons=50, dimensions=1)
nengo.Connection(stim, b, function=lambda x: x-1)
nengo.Connection(b, out_b.neurons,
function=lambda x: 1,
transform=[[-3]]*out_b.n_neurons)
nengo.Connection(out_b, output, function=lambda x: 1)
probe_input = nengo.Probe(stim)
probe_output = nengo.Probe(output, synapse=.01)
sim = nengo.Simulator(model)
sim.run(6)
import matplotlib.pyplot as plt
plt.plot(sim.trange(), sim.data[probe_input])
plt.plot(sim.trange(), sim.data[probe_output])
plt.legend(['Input', 'Output'])
plt.show()
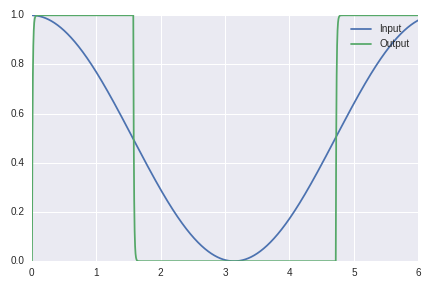
Here’s another one and look it’s perfect:
import nengo
import numpy as np
model = nengo.Network()
with model:
stim = nengo.Node(lambda x: np.cos(x)*.5+.5)
output = nengo.Node(lambda t, x: 0 if x < 0.5 else 1, size_in=1)
nengo.Connection(stim, output)
probe_input = nengo.Probe(stim)
probe_output = nengo.Probe(output, synapse=.01)
sim = nengo.Simulator(model)
sim.run(6)
import matplotlib.pyplot as plt
plt.plot(sim.trange(), sim.data[probe_input])
plt.plot(sim.trange(), sim.data[probe_output])
plt.legend(['Input', 'Output'])
plt.show()
Ok, I’m just kidding with a node you can implement anything.
1 Like
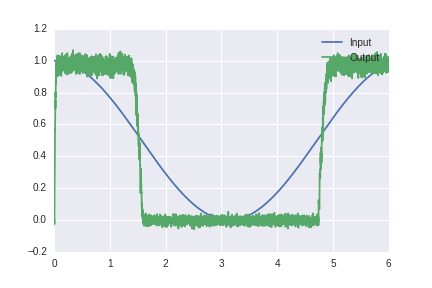
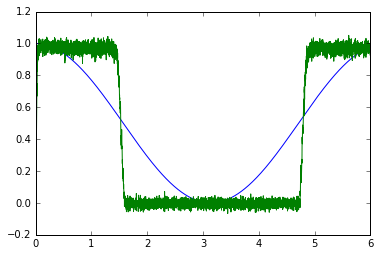
This is an actual solution with neurons:
import nengo
import numpy as np
model = nengo.Network()
with model:
stim = nengo.Node(lambda x: np.cos(x)*.5+.5)
threshold = nengo.Ensemble(
75, 1,
intercepts=nengo.dists.Exponential(0.3, 0.5, 1.),
encoders=nengo.dists.Choice([[1]]),
eval_points=nengo.dists.Uniform(.5, 1.))
nengo.Connection(stim, threshold)
output = nengo.Ensemble(75, 1)
nengo.Connection(threshold, output, function=lambda x: 0 if x < 0.5 else 1)
probe_input = nengo.Probe(stim)
probe_output = nengo.Probe(output, synapse=.01)
sim = nengo.Simulator(model)
sim.run(6)
import matplotlib.pyplot as plt
plt.plot(sim.trange(), sim.data[probe_input])
plt.plot(sim.trange(), sim.data[probe_output])
plt.legend(['Input', 'Output'])
plt.show()
1 Like
Interesting! Compressed down to one population, using 25 less neurons!Exponential more effective here than a Uniform?
Actually I though I was using the same number of neurons …
Yes, see here and here .
xchoo
August 8, 2016, 8:06pm
7
It should be noted that with the nengo presets (in the dev version of nengo), you can implement the network like so:
import nengo
import numpy as np
model = nengo.Network()
with model:
stim = nengo.Node(lambda x: np.cos(x)*.5 + .5)
threshold_val = 0.5
with nengo.presets.ThresholdingEnsembles(threshold_val):
threshold = nengo.Ensemble(75, 1)
nengo.Connection(stim, threshold)
output = nengo.Ensemble(75, 1)
nengo.Connection(threshold, output, function=lambda x: 0 if x < 0.5 else 1)
probe_input = nengo.Probe(stim)
probe_output = nengo.Probe(output, synapse=.01)
sim = nengo.Simulator(model)
sim.run(6)
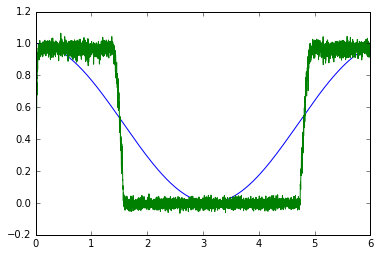
There is a slight difference in the first parameter of the Exponential distribution, but that just affects the steepness of the transition point.
Without preset:
It should also be noted that the presets are not part of any official release yet.
Great tutorial Jan! Super well put together, thanks for the link!