Thanks again for all your input!
First, I’d like to signal a bug I’ve found in nengo_extra.graphviz.net_diagram(): with the updated code you posted, I’m getting the following output graph:
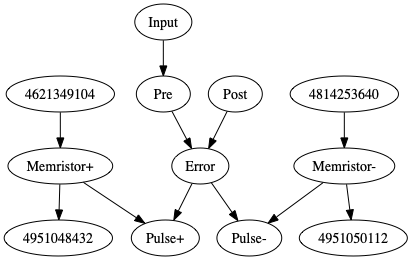
but nengo_gui renders correctly.
Returning to my problem, with this setup the input to memristor_plus and memristor_minus are the weights associated to each neuron in the pre ensemble? Or the some sort of “neural activity” (what does this term mean in this context, exactly?)?
I have checked the output of the pre = nengo.Ensemble(100, dimensions=1, label="Pre") population and it is a (100,) vector. What does each element represent, exactly?
I’m asking this because I’m trying to understand how to actually “filter” using the memristor function:
# filter the input def filter( self, t, x ): self.history.append( self.R ) # scale weights for filtering w = (self.R - self.r0) / (self.r1 - self.r0) import time time.sleep(0.01) return w * x
Until I’m sure what x actually is, I’ll have some trouble conceptualising if what I’m doing is correct or not…
Thanks again!